KryoFlux - Flux Decoder
2009-11-19
The density graphs are not quite right in Turrican 2 when dumping with KryoFlux, this is mostly due to very different resolution quantized to something CTA (SPS Disk Image Analyser) can process. CT (SPS Disk Imager) would never sample certain differences that show up with KryoFlux. This is not a big problem, and can be fixed another time.
The real problem with this game is that it violates recording rules with its sync value. It’s $9521, or 1001 0101 0010 0001.
The number of 0s before the last ‘1’ is 4. A real FDC would grow it’s cell window each time a flux reversal is not detected, making the duration of each adjacent 0 slightly longer as time progresses. This makes it unlikely to be able to sample too many 0s using any FDC, unlike when using things like KyroFlux.
It works backwards as well. If the interval between flux reversals shorten, so does the expected interval for each bitcell. This is done to try and compensate for drive speed changes during recording as well as reading. So as long as 1s are frequent enough, the correct bitcell width can be tracked.
The PLL is continuously trying to lock on to the “expected” (and continuously adjusted) cell rate. If it fails, depending on implementation it may: ignore a cell, “merge” cells, or most commonly, insert a flux reversal anyway. This is how you end up with weak bits if writing too many 0s, or other patterns that would trip the PLL.
KryoFlux uses the reference clock (in this case, 2us) to decide the size of a cell and the number of bits represented between two flux reversals. This perfectly reflects how the data was mastered, what was written for real - not how the data would be read. Therefore it can read even a thousand zeroes and would still say that it’s a thousand 0s, and not some random series of 1 and 0.
KryoFlux’s flux reversal decoder has no knowledge whatsoever on any recording format on purpose - it can decode anything. For example, you wouldn’t want to have two 1s in MFM, the distance is too short (2us dd or 1us in hd), but it’s fine in FM or C64 GCR (4us or at least 3.2us) without cells repelling. As long as you stick with tracking the cells with the reference clock, it does not matter, they’d be read perfectly.
The above pattern is already violating the rules (although it’s a don’t care for KryoFlux) but it’s been recorded with a fairly short bitcell size, probably about 1.75us. If the cell width was 2us, four 0s would be read as four 0s as discussed already, the time between the flux reversals is:
4 * 2 = 8 us
Now take a look at what happens with 1.75us
4 * 1.75 = 7 us
So, would you say this is:
3 * 0 recorded with 7/3 = 2.3333.us cells
OR
4 * 0 recorded with 7/4 = 1.75us cells
?
You could tell that 1.75 is 12.5% error, 2.3 is 16% error. In fact the first version of our flux decoder did just that. However doing so introduces aliasing for long sequences as the error very quickly diminishes to a point where it’s not possible to tell which one is the correct - but using the reference clock would give the correct number for the cells, as a real world error only effects the position of the flux reversal, not all the cells they represent.
If the decoder tracked the cellsize it would have already lowered of it’s expectation of cell size to be around the -12.5% value - but it would be unable to track sudden changes. The current one can read those without any problem.
Ideally:
- The decoder would track cellsizes.
- It would use the reference clock for questionable content.
- It would try to do post compensation to alter cells taking into account how adjacent cells react to each other according to the distance between them.
It is obviously quite some work to get this “right”, and it would still fail in some cases. The right thing to do is to have a best guess, then if it fails re-decode using different parameters.
The decoder could also highlight each cell where a decision had to be taken - effectively building a decision tree for an analyser, for example “this sequence is surely xxx, but at this point the following two bits could be either y or z, try whichever works”. The analyser then would try the most likely main route (the choices the flux decoder made) and if that fails, it would then try to traverse the decision tree finding the best solution.
If all else fails after this point (the chances are the data is unreadable... and way beyond anything that is readable by normal means anyway), the user could be still using an editor finding questionable cell sizes and manually alter the data if they are feeling lucky. :)
...but all these are in the realm of very serious time being spent on this thing. Possibly many, many months to get it right. It’d have to be a full time job.
Index Matching
Fixed index matching when the index is in a very long flux transition.
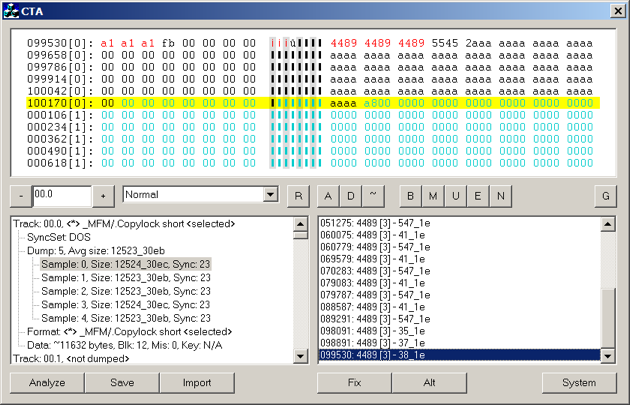
From the Turrican discussion above, you know that KryoFlux can perfectly dump tons of 0s just as they are (something a normal floppy controller cannot cope with). Now take a look at this picture of a track, dumped using KryoFlux, and shown in CTA.
The KryoFlux host software refused to accept this track as properly dumped, and got stuck retrying it. The reason was that at the position indicated by the board it could not find any reference flux reversal for index signals. All the decoders have loads of sanity checks, this is just one those.
As it happens, a very long sequence of 0s (i.e. no flux reversals whatsoever for a long period of time) start just before the end of the track and pretty much end just before data starts at the next revolution. The sequence is in fact so long, that apart from artificially generated test data, it’s been the first time we have ever seen an extended flux reversal counter representing such a long time period.
The extended sequence however has a different representation, and the decoder got confused as it did not take that into account, meaning the index matching algorithm correctly identified a lack of flux reversal where it should have been according to the index signal description.
The funny part: the game in question is called “Guardian Angel”. Nomen est omen. It surely did highlight a serious omission in the decoder. :)
![[logo]](/lib/tpl/sps/layout/header-logo.png)
![[motto]](/lib/tpl/sps/layout/header-motto.png)